Congratulations to our colleagues for constantly improving the conditions for laboratory animals.
Abstract
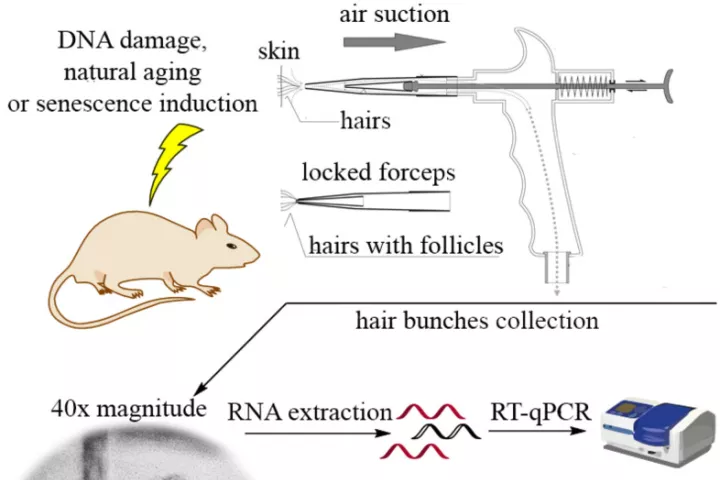
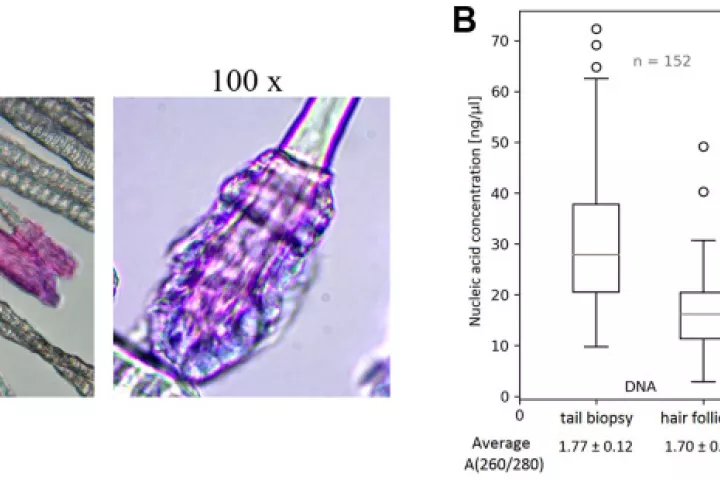
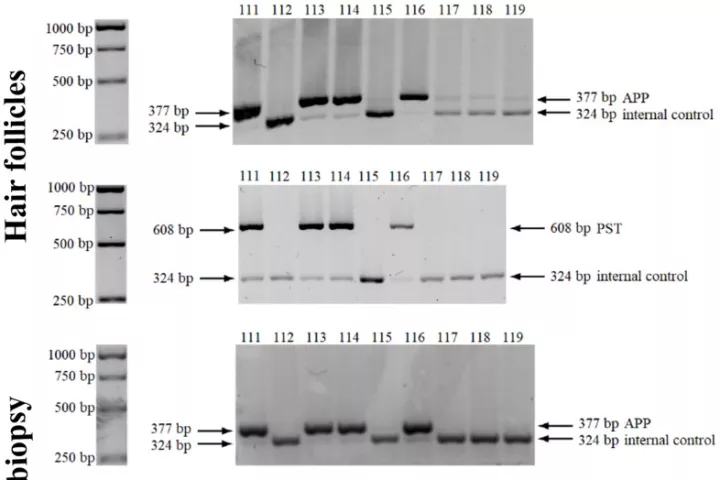
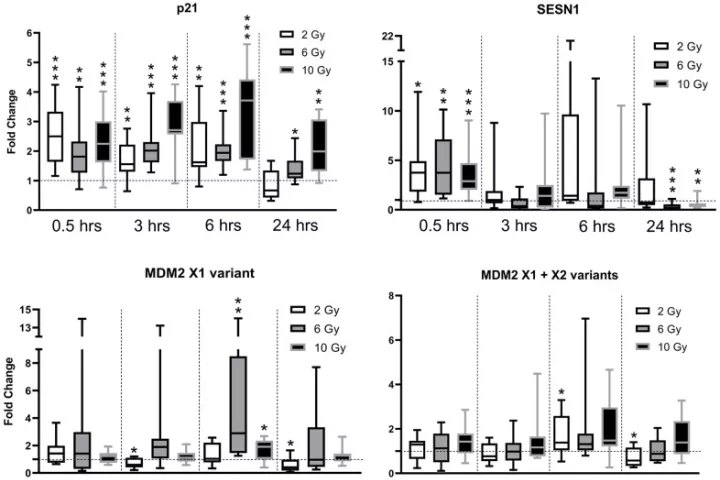
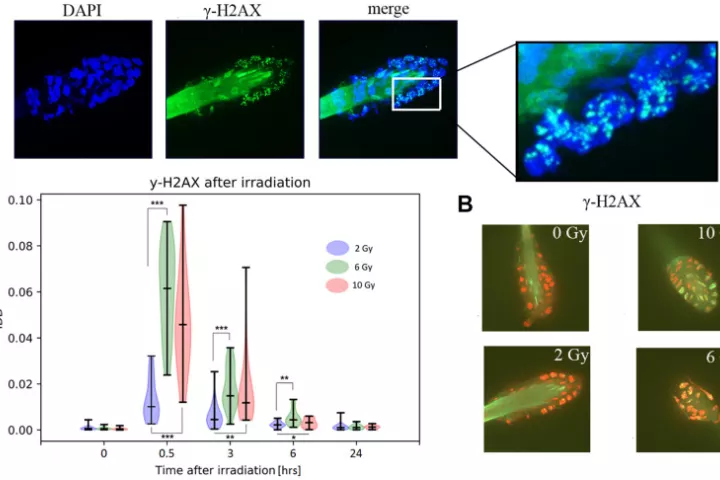
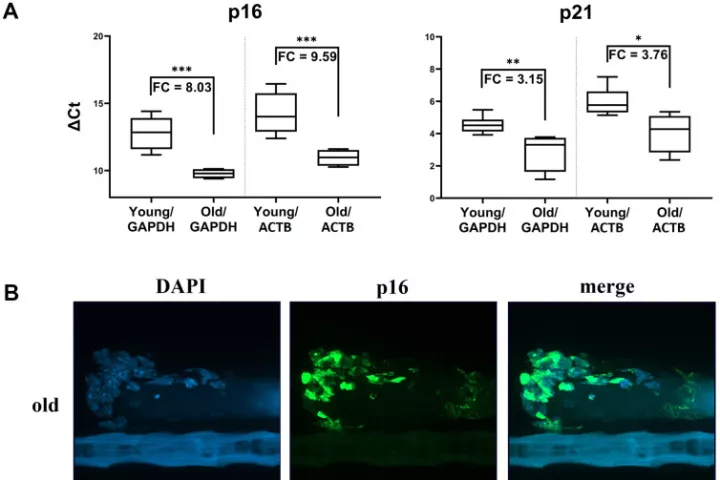
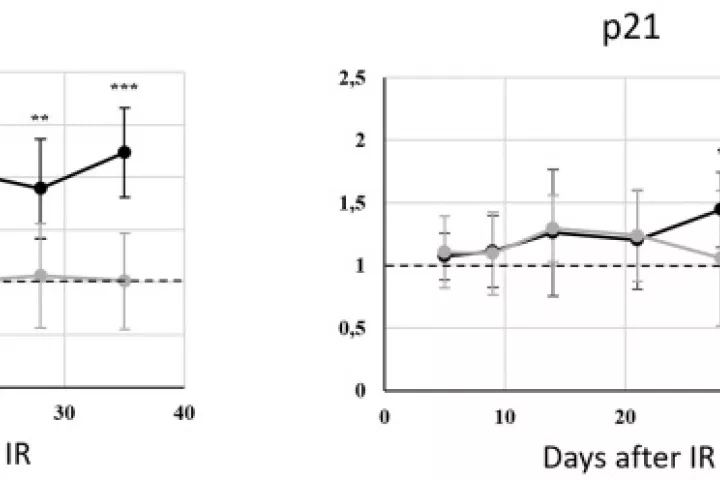
In accordance with the 3 Rs principle (to replace, reduce and refine) animal models in biomedical research, we have developed and applied a new approach for sampling and analyzing hair follicles in various experimental settings. This involves use of a convenient device for non-invasive collection of hair follicles and processing methods that provide sufficient amounts of biological material to replace stressful and painful biopsies. Moreover, the main components of hair follicles are live cells of epithelial origin, which are highly relevant for most types of malignant tumors, so they provide opportunities for studying aging-related pathologies including cancer. Here, we report the successful use of the method to obtain mouse hair follicular cells for genotyping, quantitative PCR, and quantitative immunofluorescence. We present proof of concept data demonstrating its utility for routine genotyping and monitoring changes in quality and expression levels of selected proteins in mice after gamma irradiation and during natural or experimentally induced aging. We also performed pilot translation of animal experiments to human hair follicles irradiated ex vivo. Our results highlight the value of hair follicles as biological material for convenient in vivo sampling and processing in both translational research and routine applications, with a broad range of ethical and logistic advantages over currently used biopsy-based approaches.
Keywords: DNA damage response; cellular senescence; hair follicle cells; irradiation; non-invasive sampling.
Conflict of interest statement
CONFLICTS OF INTEREST: MM, JB, MH are co-inventors on patents EP2928382A1 and US 20150297253, utilizing the method for extraction of follicular cells and device for the pursuance of this method. All authors declare they have no conflicts of interest.
The article is available HERE on Pub.Med.org.